Lung adenocarcinoma is the predominant non-small-cell lung cancer (NSCLC) subtype accounting for ~40–50% of all lung cancer cases. Lung cancer early detection by means of screening trials with low-dose CT have shown to be effective in reducing lung cancer mortality (of a ~20-30%). Yet, a considerable fraction of patients (~30–40%) with early-stage disease will experience an adverse outcome which makes the identification of prognostic biomarkers nowadays paramount.
In 2007, we reasoned that developing prognostic markers for early-stage lung adenocarcinoma was of particular relevance in order to identify the subgroup of patients who would benefit of more aggressive therapies and finally reduce lung cancer mortality. We successfully identified a 10-gene expression signature able to recognize poor prognosis lung adenocarcinoma patients even with stage I disease. Importantly, this 10-gene signature is composed by genes involved in disparate mechanisms but synergistic in cancer progression: i) cell proliferation (E2F1, E2F4, RRM2, MCM6), ii) development/stemness (HOXB7, SF3B1), cell migration and invasion (SERPINB5, NUDCD1, HSPG2), iii) and epithelial airway differentiation (SCGB3A1).
Ten years later, we moved toward the clinical application of this signature throughout the development of a robust prognostic model based on the expression of these 10 genes, by using qRT-PCR from FFPE samples, and validated in over 1,400 lung adenocarcinoma patients from different case collections and across different platforms. Very interestingly, we started to investigate the molecular characteristics of the subset at worse prognosis, and discovered that patients composing this aggressive subtype had peculiar molecular characteristics resembling those of advanced stages with high tumor mutational burden (TMB), as well as an enriched inter- and intra-tumor genetic heterogeneity (e.g., mutations, CNV, gene fusions). However, the biological processes and mechanisms driving the emergence of aggressive lung adenocarcinoma needed to be further explored.
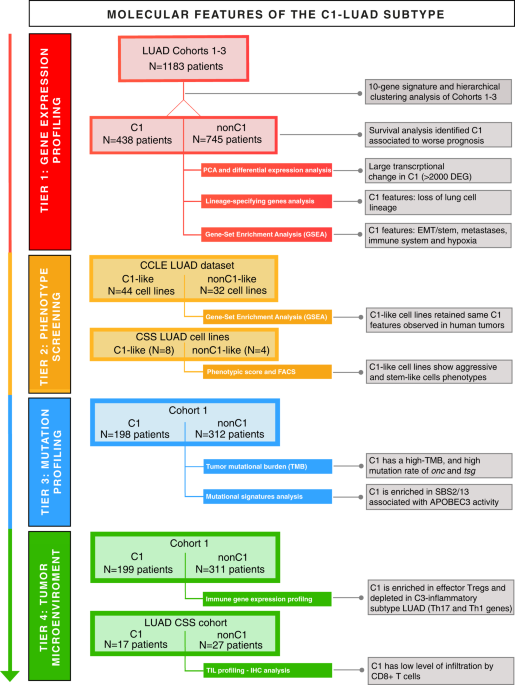
In the present study, we moved further toward the knowledge of more aggressive lung cancers by adopting a multi-tiered approach from molecular to functional characterization of lung adenocarcinoma, and by using multiple cohorts of patients (for a total of 1,227 patients) as well as lung cancer cell lines. We investigated the tumor transcriptome, the mutational and immune gene expression profiles of aggressive lung tumors (namely “C1” subtype) identified by using our 10-gene signature. We found that loss of lung cell lineage and gain of stem cell-like characteristics, along with mutator and immune evasion phenotypes, explain the aggressive behavior of the “C1” early-stage lung adenocarcinomas. Yet, using several lung adenocarcinoma cell lines we recapitulated this more aggressive subtype and functionally validated its metastatic behavior. Our data support the hypothesis that transformed lung stem/progenitor cells and/or reprogrammed epithelial cells with cancer stem cells characteristics are hallmarks of the aggressive subset of lung adenocarcinomas. Such discoveries advise alternative, more incisive, therapeutic strategies for patients with early-stage “C1” lung adenocarcinoma subtype.
We are now working on further investigations to explore the causal relationships between lung cancer molecular alterations and the acquirement of the aggressive behavior, which will pave the way for developing more effective therapeutics in lung adenocarcinoma patients.







Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in