Being located in Utrecht, the capital of organoid technology, our lab has been involved in several projects to characterize the genomes of tumor derived organoids from cancer patients. These mini-tumors represent the tumor tissue of origin and thus enables the exploration of targeted therapies in the lab without burdening the patient 1. As homologous recombination deficiency (HRD) is commonly observed in breast cancer, we wondered which breast cancer organoid would be sensitive to olaparib and niraparib, two widely used poly (ADP-ribose) polymerase inhibitors (PARPi) 2.
HRD tumors rely on more error-prone DNA repair pathways that leave behind distinct mutational scars (aka mutational signatures), such as “single base substitution signature 3” (SBS3). For the breast cancer organoids, we used SBS3 to select the most likely HRD organoids which indeed were sensitive to PARPi. Later on, more and more researchers, clinicians and cancer diagnostics programs (e.g. Hartwig Medical Foundation; HMF) became interested in a scar-based detection approach for HRD, as it was shown that twice as many HRD breast tumors could be identified compared to using HR gene driver mutations 3. Since no easily deployable HRD detection tool existed at the time, we decided to develop a Classifier of HOmologous Recombination Deficiency (CHORD - also in reference to our love for music).
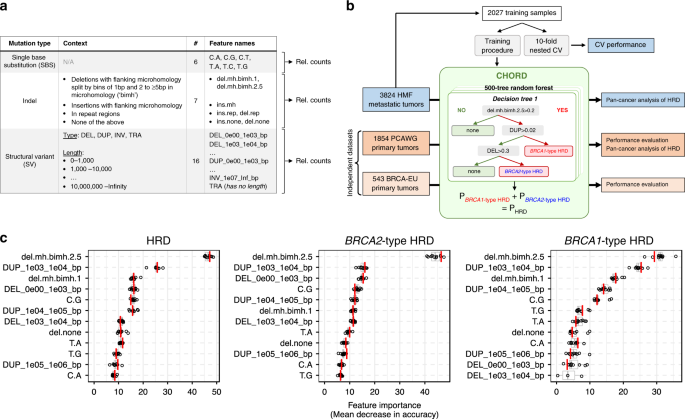
Conceptually, we had two main objectives: i) rely on simple, but robust, mutation features and ii) develop a tissue-agnostic HRD biomarker. Seven years after the mutational signature hallmark paper 4, a consensus tool for signature extraction is still missing, partly because signature extraction is complex. Recommendations for optimal extraction procedures have been discussed such as a priori selection of tissue specific signatures to reduce overfitting 5,6. However, such requirements are far from ideal in clinical workflows, e.g. because tumor tissue-of-origin information is often inaccurate or missing (e.g. CUP cancer patients). In our paper, we show that CHORD can accurately predict HRD without the use of mutational signatures. Moreover, we find that SBS3 but also point mutations in general are actually of little value for predicting HRD (in agreement with others 3,6). The second objective relates to applicability of CHORD to (future) WGS diagnostic workflows. Since we found that HRD is basically in every cancer type, HRD detection must be independent of tissue related mutation processes. We have indeed shown that CHORD can be applied to any cancer type and thus would not hamper clinical practice in routine diagnostics.
Using the predictions from CHORD, we revealed that HRD is most frequent in ovarian and breast cancer, but also prevalent in pancreatic and prostate cancer (Figure 1). We have also made several findings with clinical implications. First, we found biallelic HRD inactivation solely by somatic events in ~25% of HRD patients, which would remain undetected if only germline mutation screening is performed which is currently common practice in many labs. Secondly, we found that deep deletions of HRD driver genes occur in 10% of HRD patients pan-cancer which would also be missed with ‘traditional’ mutation screening since there are no point mutations to detect! This is especially common in prostate cancer where 33% of the tumors have a complete loss of an HRD driver gene. Lastly, although PARPi are in general only approved for patients with breast, ovarian, prostate and pancreatic cancer, we have shown that ~15% of HRD patients have other cancer types (Figure 1).
Most notably, CHORD is already part of the HMF patient diagnostic report for over 2 years, which has been vital for the interpretation of mutations in HR genes as well as treatment decisions with PARPi (e.g. within the Dutch DRUP study 7). This has enabled more and more patients to benefit from HRD targeted therapies, and we believe that CHORD will continue to have such impact in the coming future.

Figure 1: Frequencies of HRD pan-cancer (left) and across different cancer types (right).
References
- Drost, J. & Clevers, H. Organoids in cancer research. Nat. Rev. Cancer 18, 407–418 (2018).
- Sachs, N. et al. A Living Biobank of Breast Cancer Organoids Captures Disease Heterogeneity. Cell 172, 373-386.e10 (2018).
- Davies, H. et al. HRDetect is a predictor of BRCA1 and BRCA2 deficiency based on mutational signatures. Nat. Med. 23, 517–525 (2017).
- Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
- Maura, F. et al. A practical guide for mutational signature analysis in hematological malignancies. Nat. Commun. 10, 1–12 (2019).
- Degasperi, A. et al. A practical framework and online tool for mutational signature analyses show intertissue variation and driver dependencies. Nat. Cancer 1, 249–263 (2020).
- van der Velden, D. L. et al. The Drug Rediscovery protocol facilitates the expanded use of existing anticancer drugs. Nature 574, 127–131 (2019).





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in