Acute myeloid leukemia (AML) is an often fatal blood cancer. Several novel drugs have been approved for specific AML subgroups, but most patients are still treated with conventional chemotherapy. Chemotherapy effectively kills leukemic cells and thereby leads to remissions in many patients, yet a large proportion go on to relapse. Relapse is typically therapy resistant, and a major cause of death in AML.
To investigate molecular mechanisms leading to therapy resistance at relapse, we had previously compared gene expression patterns between diagnosis and relapse of AML. The idea was that genes whose expression differs between these two disease stages in a majority of patients (i.e., non-randomly) are likely to contribute to their specific properties, most importantly, their different drug responsiveness.

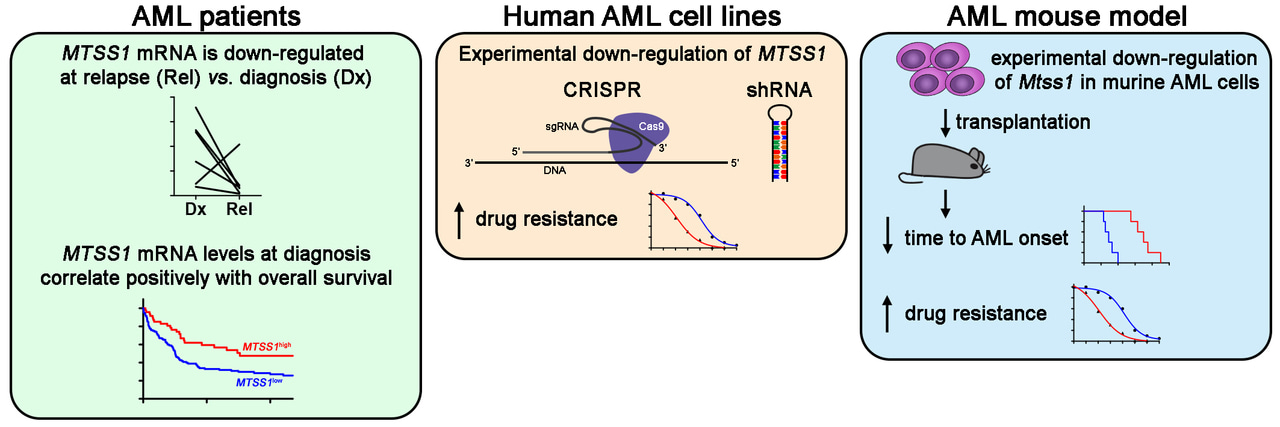
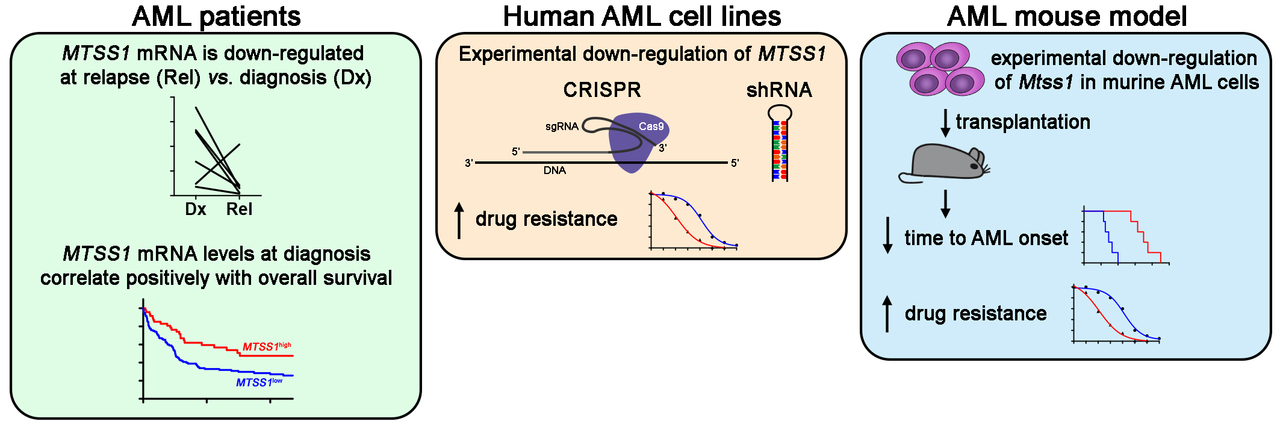
One of the genes that was identified in this manner was metastasis suppressor 1 (MTSS1). MTSS1 was down-regulated at relapse of AML. Low MTSS1 expression was already known to contribute to the formation and progression of some other tumors. We therefore investigated whether it would also have a role in therapy resistance and aggressiveness of AML.
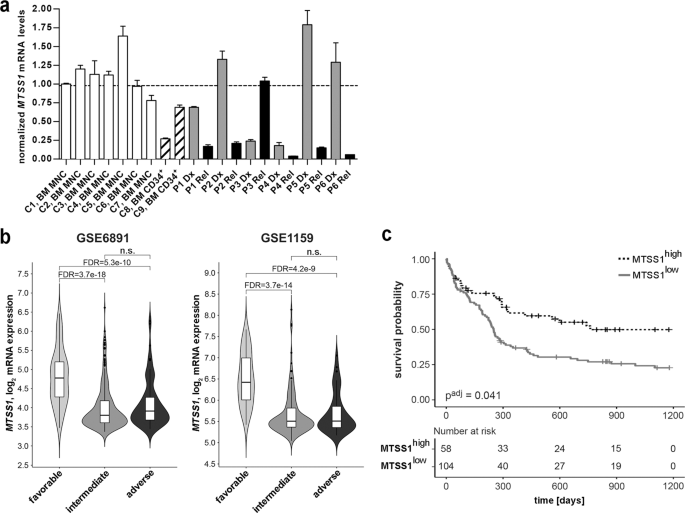
First, we tested whether MTSS1 expression at diagnosis of AML was related to overall survival of patients. For this purpose, we used gene expression data that are freely accessible through the database Gene Expression Omnibus – a really cool resource that allows one to generate additional support for a hypothesis (or to dismiss it) in short time and without any pipetting. Indeed, AML patients with low MTSS1 expression at diagnosis had significantly worse outcome than those with high MTSS1 expression.
Next, we used two different methods to reduce or eliminate the expression of MTSS1 in three different AML cell lines – the ”modern“, Nobel prize-awarded CRISPR method, and the more ”old-fashioned“ shRNA method. In our context, old-fashioned shRNA had the advantage that it leads to a reduction, rather than an elimination, of gene expression – this better reflects the situation seen in human AML patients. Nevertheless, with both methods we found that cells with low MTSS1 expression were killed less effectively by AML chemotherapy than those with high expression.
We went on to ask whether we would be able to identify any drugs that would work better on cells with low than with high MTSS1 expression, and might serve as therapeutic alternatives for such patients. Among over 100 different anti-cancer drugs that were tested in a robot-assisted screen, none fulfilled this hope. Rather, MTSS1 down-regulation increased resistance towards nine additional drugs, suggesting a more general role of the reduction of MTSS1 expression in drug resistance.

To confirm the role of MTSS1 in a living organism, we used a mouse model of AML. Experimental reduction of Mtss1 expression led to a huge acceleration of AML development, and also augmented other indicators of disease aggressiveness. Moreover, in leukemic cells isolated from the mice, reduced Mtss1 expression increased chemotherapy resistance.

Together, these data confirm the hypothesis we had raised based on the expression patterns of MTSS1 in human patients, namely, that reduced expression of MTSS1 contributes to aggressiveness and therapy resistance of AML.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in